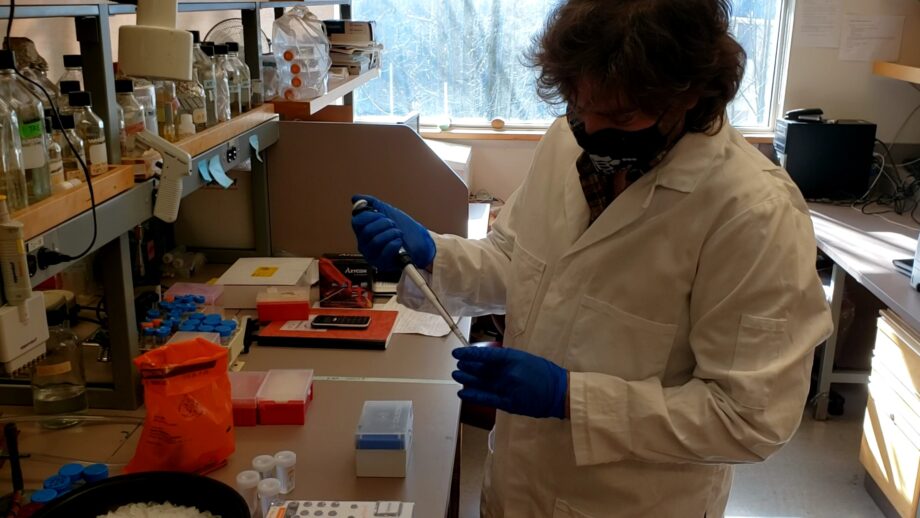
Sequencing the Crisis: How genomics morphed from a COVID-19 research tool to a critical part of the pandemic response
By Mary Gooderham
The word that came from public-health officials in the United Kingdom on Dec. 20, 2020, was an unwelcome—yet not altogether unexpected—challenge for the genomics community in Canada.
A variant of the SARS-CoV-2 virus that causes COVID-19 had been found with mutations in its spike protein that made it much more transmissible than the original strain identified in Wuhan, China. First detected in a sample taken in southeast England in September, it was quickly spreading among the population there.
News of this new strain—it was known as lineage B.1.1.7 and would come to be called the Alpha or U.K. variant—spread among members of the Canadian COVID-19 Genomics Network, or CanCOGeN. The group had been launched in April 2020 with a mandate to sequence the genomes of the COVID-19 virus and people infected with it in Canada. Here was a new purpose, urgency, and frankly, point to the past months of exhaustive surveillance work.
“That was absolutely a defining moment,” recalls Dr. Catalina Lopez-Correa, CanCOGeN’s executive director. “The presence of the variants of concern, as we came to know the Alpha and other variants, was a game-changer.”
The announcement regarding the Alpha strain would cancel everyone’s plans for the holidays. If anyone was looking forward to a break right then it was Lopez-Correa, who had arrived in Vancouver just two days earlier from her native Colombia, where she’d been working remotely for CanCOGeN for six months. Any notion of spending her time in quarantine “unplugging and resting” went by the wayside. There were daily conversations with Health Canada, the National Microbiology Lab (NML) in Winnipeg and provincial public-health labs across the country trying to track down the variant.
“It was very intense,” says Lopez-Correa, a medical doctor and genomicist who trained and worked in the burgeoning field in Colombia, France, Belgium, England, Iceland and the United States before coming in 2008 to Canada, where she served as chief scientific officer of Génome Québec and Genome BC. She went back to Colombia in November 2019 to head an innovation hub there, and with the outbreak of the virus, turned to helping ramp up the country’s COVID-19 treatment and testing program. Then her attention shifted to an impressive Canadian genome-sequencing initiative taking shape under the leadership of Genome Canada.
The effort—a consortium of public-health authorities and their health-care partners, academia, industry, hospitals, research institutes and large-scale sequencing centres—had begun just as COVID-19 was bringing the world to its knees. Indeed, even before the declaration of the pandemic by the World Health Organization on March 11, 2020, Genome Canada heard from two Canadian university labs looking for funding to do some genomic research on the virus, recalls Dr. Rob Annan, the organization’s president and CEO.
“We co-ordinated a couple of quick conversations with them and then realized that we should put together a plan and a funding proposal for sequencing the virus,” remembers Annan, a biochemist and geneticist who’s spent most of his career working in science policy.
While Genome Canada is a national organization, the initiative “really was a grassroots effort,” Annan says, adding that people quickly looked to help. “The whole world was shocked by the arrival of this pandemic.”
“Scientists are scientists, and when you have a hammer, you look for nails. And genomics is a powerful hammer,” he explains. “So, the genomics community said, ‘Okay, we need to start sequencing the virus.’”
The question was how to make this into a national program, Annan says. “We were probably a little naive, frankly, about just how big a challenge we would face in that effort. But it was started by the scientists themselves, who said, ‘We’re going to start sequencing and monitoring this virus. And we will leave it to you guys to figure out how to actually knit this thing together.”
Within five days, Genome Canada had put a proposal into the hands of its partners at Innovation, Science and Economic Development Canada (ISED). “We needed to put a plan in place, and we would use that to bring people into the fold … Then it was building from there,” Annan says.
The proposal outlined a $40-million project focused on genomics: the science of deciphering and understanding the genetic information of organisms, including humans, animals, plants and microorganisms, encoded in DNA and related molecules. CanCOGeN would sequence the genomes of both the virus and people exposed or affected by it.
VirusSeq and HostSeq
The viral-surveillance program, costing $20 million and eventually known as VirusSeq, would involve federal and provincial public-health labs and academic institutions across the country, sequencing the virus in up to 150,000 positive COVID-19 cases. The SARS-CoV-2 genome consists of nearly 30,000 RNA “letters”, a relatively small sequence. Sequencing individual genomes from viral samples across time and space would track how the virus was changing and spreading, providing data that could help with viral detection, for example, or the effectiveness of treatments and vaccines.
“It was useful for the public-health response,” explains Dr. Guillaume Bourque, an associate professor of human genetics at McGill University. He says that when the project started, “everybody was ecstatic, myself included.” It was “an amazing opportunity” that something scientists had been working on for years could be applied to a problem on the front page, says Bourque, who is also director of bioinformatics at the McGill Genome Centre and director of the Canadian Centre for Computational Genomics.
Another $20 million would go toward sequencing the genomes of 10,000 people exposed or affected by COVID-19. Called HostSeq, it would look at possible genetic factors behind the variability in patient outcomes—for example, why disease severity varied greatly among people. Human DNA includes 3 billion base pairs (around 20,000 genes), bringing much greater complexity to its sequencing. Co-morbidity and environmental factors, as well viral mutations, were also elements to consider.
“The host side was always a longer-term game, the idea of how we can design drugs or vaccines as we go forward,” explains Dr. Naveed Aziz, a geneticist who is CEO of CGEn, Canada’s national platform for genome sequencing and analysis. It oversees the three major facilities in Montreal, Toronto and Vancouver that would do the host sequencing.
Dr. Lisa Strug, director of the University of Toronto’s Data Sciences Institute and a senior scientist at the Hospital for Sick Children, had heard from a number of colleagues in fields such as immunology and infectious diseases looking to sequence small cohorts of people infected with the virus. “Each research group had a different question,” says Strug, a data scientist in the field of complex human traits. Laying the groundwork for a national COVID-19 human genomics databank would be “a humongous win,” she says, providing a resource that could support a wide range of research.
A network effect
The creation of CanCOGeN was announced on April 23, 2020, under the federal government’s coronavirus research and medical countermeasures investment. Annan points to “phenomenal leadership” by federal departments such as ISED, the Public Health Agency of Canada (PHAC) and Health Canada, where Deputy Ministers convened various leaders’ round-tables on COVID-19 to create policy synergies and co-ordinate the different components of the federal COVID response.
There was a lot of work to be done after the initiative was launched by Genome Canada and the six regional genome centres, Annan notes. This included “hard infrastructure” (the machines and other equipment needed to do the sequencing), and “soft infrastructure”, setting up sharing protocols and making sure data analysis was done uniformly, governed by appropriate privacy and security measures. “Those tend to work provincially,” he says, and Canada’s decentralized healthcare system was already presenting challenges. “There was a network effect starting to happen to bring the groups together” into a national consortium, he says, which included the NML, academic and provincial public-health labs.
Annan comments that PHAC had been created following the SARS outbreak that hit Canada in the early 2000s to build out a national public-health response, “but we didn’t make the kind of sustained investments that would allow us to be fully ready for the next one.” There certainly wasn’t the scale of investment necessary to build and maintain a national genomics surveillance system. CanCOGeN was forging a largely new structure to respond to SARS-CoV-2.
Guillaume Poliquin, interim vice president of the NML, says one legacy of the first SARS crisis was that his lab was able to develop the first functional test for SARS-CoV-2 in mid-January, five days after Chinese researchers published its sequence online. “The reason we did that is because with SARS 1, we had developed exploratory diagnostic tests that would be able to pick up new coronaviruses if they emerged,” he says. “So that capability was there, and it was used.”
Poliquin, a pediatrician and infectious disease physician with a specialty in medical microbiology, says that before the COVID-19 pandemic, there was a lot of work being done to get the NML and the Canadian Public Health Laboratory Network ready for the “onboarding” of genomic technology, as the “next real sea change in the way that diagnostics and particularly surveillance was being done in Canada.” Like any major transformation, says Poliquin, “it was not an overnight type of process.”
COVID-19 accelerated these plans, and indeed things needed to happen overnight, to put it mildly. CanCOGeN was conceived as a surveillance and research initiative, Poliquin says, with implications for vaccines and outbreak investigations. But the “biological significance” of virus variants that would come along, like the Alpha and soon the Delta strain, meant that the genomes “needed to be looked at with a higher degree of fidelity and much faster turnaround time than what was initially envisioned,” Poliquin says.
Meanwhile, provincial labs were creating internal structures and capabilities to deal with the crisis. “Contributing to this big national effort was not top of their priority list,” Annan says, which left CanCOGeN “cajoling the provinces to participate.” Additionally, the labs needed funding and other resources “just to create the bandwidth to feed up into that national system.”
Key Takeaway: Canada—like many others—didn’t fund national pandemic preparedness in a sustained manner, and we saw the dire consequences.
An evolving need for sequencing
Out in the field, many things were happening in parallel. Dr. Natalie Prystajecky, an environmental microbiologist who is program head of the British Columbia Centre for Disease Control (BCCDC) public-health lab and clinical associate professor at University of British Columbia, notes that the first genomic sequencing of the virus began as a research project. At the end of January 2020, three months before CanCOGeN even launched, the BCCDC received a rapid response grant from Genome BC, a Genome Canada provincial counterpart, to develop a method to sequence the COVID-19 genome.
“Within days we had our first positive case,” Prystajecky recalls. Medical-health officers and infection-control practitioners on the frontline were suddenly asking the scientists to sequence the outbreak and tell them what was happening with patients experiencing different symptoms of the virus.
“It became very clear that this was not just a fun research project, that it had a function in the pandemic response,” Prystajecky says, noting that the lab was then sequencing a few cases a day. When support came from CanCOGeN, “we realized that we could transition this from a research project, with research staff, to a public-health service, with our staff.”
By the end of August 2020, the lab was producing 200 to 300 viral genomes a week, Prystajecky says, covering about 20% of cases. “I felt like we were really staying on top of things.” But there were new waves and the purpose of sequencing changed over time, she says, with people asking questions like, “Are there reinfections?”
Things morphed further with the Alpha variant, “because it became clear that sequencing was more than just responding to outbreaks or asking questions about reinfections,” Prystajecky says. “We found it in BC; we knew that we were sequencing a traveller from the U.K.”
The task then became trying to find and respond to variants of concern in the province, she says. The lab was sequencing 1,000 or more samples a week, up to 100% of cases. The reason for sequencing continued to evolve; for example, the data are used today to study vaccine efficacy and understand whether there’s something different among the cases of vaccine-breakthrough infections, she says. In late September 2021, the lab was sequencing 4,000 viral genomes a week, about 80% of cases. “That’s a long cry from where we were when this project started, when we did 24 in a week, and we thought we were great.”
Through it all, the lab has struggled with quality control, burnout and frustration with pressure from CanCOGeN to contribute to the national effort, Prystajecky says. “They had rose-coloured glasses about what could be accomplished. We were in the trenches doing the work, and sometimes not always feeling as appreciated about how much work we had to do to get things done well.”
Key Takeaway: Canada’s response efforts brought to light real challenges we face due to the federated nature of our health system.
A response 20 years in the making
While Genome Canada’s response to the pandemic was rapid, Annan says, it was 20 years in the making. Since its creation in 2000, the not-for-profit organization had received some $2 billion in federal investment to support genomics research by the academic community and industry in areas such as cancer, rare diseases, agriculture, forestry and climate change. “All that investment allowed us to have the capacity to be able to mobilize very quickly and to pivot,” he says.
In CanCOGeN’s early days, Genome Canada “bootstrapped” its activities with funds from other projects and worked to build links with actors across the country. When Lopez-Correa first saw a job posting in the spring of 2020 for someone to co-ordinate the initiative, she thought that CanCOGeN “had the potential to really change the world” by genomically tracking the evolution of the virus, despite other public-health concerns.
“During the early days, people were worried about protection, masks, testing,” she remembers. Here was something much different. “You’re in a pandemic, and you’re thinking that you’re going to invest $40 million in sequencing the virus, and sequencing the human, and putting them together under the same umbrella?”
Lopez-Correa took up her position as executive director of CanCOGeN in June 2020, as funding agreements and governance structures were coming into place. Like many people caught out by pandemic travel bans, she remained in Colombia, joining video meetings from the living room of a cottage in the countryside outside of Medellín, the discussions punctuated on her side by the sounds of exotic birds and occasional tropical storms.
Much of the early talk focused on the costs and technical points of sequencing, she says, “but sequencing was not the endpoint. The real goal was how we were going to use that information to inform public-health and policy decisions.” That had already occasionally been happening, for instance with the very early sequencing showing that most COVID-19 viral lineages in B.C. were coming from the United States, Lopez-Correa points out. “Genomics was one of the data points that led to the difficult decision of closing the border with the U.S.”
Putting together this massive genome sequencing and analysis effort in the middle of a pandemic has been likened to building an airplane while flying it. Decisions had to be made at lightning speed. For example, provincial labs needed funds to do their work, but they varied considerably in how much money they requested to sequence each viral genome. CanCOGeN’s formula averaged the amounts and took into consideration the volume of cases.
“We wanted that plane to be flying—fast,” Lopez-Correa says, noting that reaching an agreement on such issues might typically have taken several months.
“There was no time to build consensus,” she says. “The provincial labs were happy because the process was transparent, it was clear, it was open. They got their money fast, they could start sequencing and building the capacity. They could start buying their equipment, hiring people and building their labs.” In some cases, as in the Atlantic region and the Prairies, this all happened from scratch, but labs across the country had challenges. “They were spread super thin, they were stressed to the max. And here we come with money to help them, but also all these requirements and all these questions. It was very, very difficult.”
Key Takeaway: The idea for CanCOGeN came together at lightning speed as a bottom-up initiative with many moving parts. And it worked.
A massive scale-up
Viral genomic sequencing “had never been done at that scale,” Lopez-Correa acknowledges. Public-health labs also typically work with sensitive information, behind closed doors. “With CanCOGeN our goal was, ‘Let’s share the information. Let’s really make this an open game and talk to each other and collaborate.’ While provincial labs welcomed new support, there were bumps along the way to navigate the new open-science, open-data approach.”
Prystajecky comments that CanCOGeN at its inception should have included representation at its senior level from the public-health labs. “That’s actually why I jumped in,” she says, offering her perspective on its implementation committees. “Provincial public health is different than federal public health. We have different functions, different mandates, different views of the world.”
Prystajecky says there were “a lot of disgruntled feelings about people telling us, ‘This is how it shall be done. And this is what it costs to sequence.’” Having the provinces involved in the planning for CanCOGeN “would have made it go a little bit slower,” she allows, but it would have reduced some of the barriers and stumbling blocks, while bringing “a better sense of buy-in.”
Viral surveillance meanwhile was beginning to happen around the world. For instance, the U.K. “was very quick out of the gate with a big national initiative,” Annan notes. Many countries were developing some form of genomic surveillance and tracking program. There was co-ordination globally among scientists to share the resulting data publicly in international databases, “because the virus obviously doesn’t care about national borders,” Annan comments.
There was less openness in Canada, notes Annan, who finds that work remains to be done on the links between the academic and public-health systems here. “In some ways, there’s even a culture clash, I would say, between the two,” he remarks. The public-health system generates data for specific purposes, like border-control measures, while academics want to get data out into the public, “and that way everyone can be working on it and figuring things out. Those two attitudes are not always well-aligned, especially in the middle of a crisis.”
Prystajecky acknowledges that the public health-academia interaction “is still really awkward … but you know, it’s something we’re trying to learn to do.” She explains that microbiologists in the world’s public-health labs “have been testing millions of samples a year for decades,” and no one paid much attention. “Academics wanting to work together, to access our data, to make suggestions on how we should do our work, that is new to us. And it’s not a natural union.”
Bourque says the first few months of scaling up and sequencing the viral genomes “were extremely exciting and satisfying,” bringing together genome researchers and public-health labs to be able to characterize the virus. “All of us were working as much as was needed, over the weekend and at night, because … by working together, we could answer questions of immediate public-health impact,” he says. “Where progress was much slower than I anticipated was in the actual data-sharing out of the provincial public-health labs.”
Lopez-Correa says the effort to address data-sharing over time included helping the provincial labs hire staff and automate their systems, as well as trying to assuage privacy concerns over the use of patient “metadata”, such as the sample collection date, location and other details that made the sequence more useful.
Key Takeaway: There is a culture clash between public health and academia. It’s natural and understandable—and we need to work through it.
A collage of data-sharing barriers
CanCOGeN issued a memo in October 2020 noting that metadata did not constitute personal information and should be shared. Three months later, the VirusSeq ethics and governance committee produced a long point-form list of impediments “real or perceived” to releasing it. “We were trying to pin down the barriers to try and find solutions for each of those barriers. We ended up with almost a mosaic, or a collage. Every province had a different challenge,” allows Lopez-Correa, who believes that at the end of the day, “provincial labs understand the value of data sharing but face significant impediments—particularly in the time of a crisis—and we had to find ways to work around them.”
Dr. Yann Joly, a lawyer specializing in health law and bioethics who is an associate professor in the faculty of medicine at McGill University, says the memo and list of impediments prompted several provinces to sign an agreement promoting data sharing across the consortium. Another important step had been setting up a VirusSeq ethics and governance committee, a small group of key people who “could really tell each other the real thing,” says Joly, the committee’s chair. “We had a lot of deep discussions, and it made it really easy for me to identify what some of the key problems were,” such as the fact that the labs were invoking data privacy as a reason for not communicating data.
Annan says the data-sharing issue especially came into focus when the more transmissible Alpha variant was identified by the U.K. in December 2020. “We needed to change our tools a little bit,” he says, to try to catch such variants of concern. This reinforced the importance of having a community-led sequencing effort by partners who were open with their results, shared their data and worked collaboratively across provincial, national and international borders.”
“The variant was changing the dynamic of the pandemic,” says Bourque, who calls the announcement from the U.K. “a big eureka moment.” Countries adapted their public-health policies; for example, Canada immediately stopped flights coming from the U.K. Bourque says there was also a realization that “we needed to be sequencing in real time,” where previously there had been more of a “passive mode” and a “retrospective analysis” of the genome data. “We realized this is not just a research project. This is part of the public-health response to the pandemic, and we had to kick into second gear with the workflow.”
Poliquin says that “CanCOGeN, whether they wanted to or not, had to move very quickly, and I think successfully, from a surveillance, primarily research-focused, endeavour into a service-delivery model in the span of weeks, which I think took everyone a bit by surprise.”
Joly feels that data-sharing “started to change a little bit for the better” after the Alpha variant was discovered, noting that until that point “we were not a well-oiled machine,” while the U.K. was “really fast in finding the right way to respond to things. For us, to set up the machine took time.”
In early 2021, the VirusSeq Data Portal was created to share SARS-CoV-2 genomic data in Canada and to harmonize, validate and automate the submission of all Canadian SARS-CoV-2 sequences and “non-personal metadata” to international databases. Bourque, who became research lead of the portal, says the key was to organize the data in a way that could be useful for public health and the scientific community both in Canada and globally. He notes that the portal has been “only partially successful” in this, and today just one quarter of the total genomes that have been sequenced in Canada are deposited in it. Bourque notes that lab co-operation remains fragmented. “We have to have conversations independently with each of the provinces to try to understand, ‘What’s the bottleneck?’”
On the HostSeq side, it proved difficult to link each of the sequenced viruses with the patients who had been infected with them. Aziz says that sequencing host genomes “needed the specific virus and the specific patient to be connected.” This was hampered by differing lab policies “and they just didn’t have the resources.” Given these limitations, HostSeq instead worked with clinician researchers who could bring together viral genomes with blood samples collected from consenting COVID-19 patients. Aziz hopes this “plan B” will mean there is a viral sequence to match up with about 40% of the host sequences.
Strug says that for HostSeq, “every headache we’ve had, the whole way through, is about data sharing, obtaining data from different institutions, organizations, provincial bodies.” This was not entirely unexpected, she says, but “the extent to which it inhibits us from doing the sorts of things that are much easier to do in other countries is staggering.” Strug would like to see a “national office of data sharing that solves this problem.”
Key Takeaway: Data sharing is a challenge with myriad elements and many players. We continue to struggle and adjust as we overcome the obstacles before us.
A shift of culture and resources
Joly feels that data sharing will improve, given improved political will among the provinces. “It’s a shift of culture for them, which is why it takes time and it takes reinforcement, but I do think that that we can get there.” He says that today, “the provinces feel a little bit more pressure and feel like it’s becoming a little bit more accepted as being part of their job, which it wasn’t at all when we started this.” He notes however that “the data sets that we get are sometimes quite incomplete in terms of the metadata that’s accompanying the genomic sequence. And yes, sometimes things are still just too slow.”
A paper published in Nature Biotechnology in August 2021 notes that Canada had a lag time of 88 days in submitting SARS-CoV-2 genome sequences to the Global Initiative on Sharing All Influenza Data (GISAID), a number five times greater than the U.K., at 16 days.
Poliquin says one reason for the delay in Canada’s contributions to GSAID is that the country is currently sending more recent data there. “Those are the sequences we want to migrate first to the international database, because they’re the ones that are most relevant to what’s happening now” Canada sends more historical information to GISAID “out of completeness,” hence the long lag time noted in the Nature Biotechnology paper. Such information “is important for trends, but it gets stale-dated very quickly,” says Poliquin, who points out that Canada, with 0.5% of the global population, has deposited 3.5% of the total sequences in GISAID. “We have punched far above our weight.”
Poliquin says solving the challenges around the sharing and accuracy of the metadata is a public policy issue “rooted in a lot of politics and history and different comfort levels,” but especially requires enhancing lab staffing. “That’s a lot harder to do in the middle of pandemic, because everyone’s busy and there’s only so many people around.”
He says that in December, “when it became clear that we need to move into an operational response,” the NML developed a “genomics liaison technical officer” or “GLTO” program to hire people with appropriate skillsets and embed them at labs doing sequencing. This provided additional capacity to handle the data and transmit it in a timely manner. GLTOs (which he pronounces “gelatos”) continue to be brought in, Poliquin says, “because the need has kept growing.” Labs with GLTOs have improved the timeliness of their data sharing, and similar improvements have happened on the process side. “We hired a systems engineer to do a walkthrough of all the systems and how they worked, so that we could identify the bottlenecks and see what types of resources we needed to apply to smooth out the pipeline.”
Poliquin notes that the U.K. has been lauded for its success in data sharing; its system is built on the National Health Service. “All of their health systems are the same, so they all speak to one another,” he says. “We are working to make a patchwork of systems work together.”
Annan considers Canada’s data-sharing challenges a reflection of deeper issues in a health system with “too many jurisdictions and decision-makers, but also a culture that prioritizes risk-minimization.” The federal government has launched a pan-Canadian Health Data Strategy to address the challenges in collecting, sharing and using health data, he notes. “Hopefully we can use this as momentum.”
Poliquin cautions that “any new technology that’s applied at scale brings about questions,” such as how people’s personal information is kept safe, and whether a combination of data might make it become identifiable. “One of the common themes has been, you need to get the sequences published and out in the public domain so that researchers can access them, researchers can make important discoveries with them,” he says. “But as holders of a lot of public-health information, we have to be great stewards of that information.”
Lopez-Correa feels that CanCOGeN should have encouraged more data sharing “from the get-go”, perhaps through different funding models. In the U.K., for example, labs are reimbursed for sequencing genomes only when the sequence is shared in an international database. A lack of sharing means that what’s going on in one province doesn’t get communicated beyond its borders, she says. “The virus is moving around the world freely, and yet we’re putting all this data in silos,” Lopez-Correa says.
Key Takeaway: The Canadian scientific community is resilient, adaptable and can learn to move at speeds hereto unthinkable.
A long list of accomplishments
Annan says that despite issues with data sharing, the building of genomics capacity and infrastructure allowed for CanCOGeN to provide important, usable information on virus spread and variants of concern to inform public-health decision making. This meant standardizing approaches, sharing best practices, collaborating on issues like governance “and building a high-quality national cohort of patient data that will be important for many years to come.”
For Joly, a major accomplishment of CanCOGeN was “the way that people came together: people that were not used to collaborating, people that might have very different views.” He saw industry, public health, academic researchers and government people trying to “settle their differences and work in the same direction.” Those from closed cultures “opened up a little bit,” while typically bureaucratic agencies with heavy administrative procedures “managed to really tone it down” and relax requirements like reporting standards.
“Things happened much more quickly when decisions needed to be taken and the money needed to flow,” Joly says. “If we can build on that for the next time around, in a more co-ordinated plan, this can really become interesting.”
Aziz says some 7,000 of the host genomes have been sequenced since work on collecting them began in April 2021. A total of 14,000 host samples have been gathered to date, while only 10,000 had been anticipated, he says. “Now the challenge is how do we fund the sequencing and data generation of the other 4,000.”
HostSeq is a “project of projects,” with the data available to principal investigators to use. Global researchers are especially interested in exchanging data with Canada, Aziz says, particularly those with homogenous populations. “The diversity in the Canadian population is an absolute goldmine.”
Meanwhile, on the VirusSeq side, CanCOGeN passed the benchmark of sequencing 150,000 viral samples this past summer. On Sept. 22, 2021, there were more than 210,000 virus genomes sequenced in Canada and growing, Poliquin says.
Genomics comes out of the lab
The COVID-19 crisis has “taken genomics out of the lab,” comments Lopez-Correa, who is passionate about science communications. She worked since 2008 in Canada to educate the public about the importance of genomics in areas such as forestry, agriculture and health. “Suddenly, with this pandemic, we have a unique way for everybody to understand genomics,” she says. “It’s now part of everyday life; people know why it’s important.”
Prystajecky agrees that it’s become “a hot topic and front of people’s minds. They are now aware of microbial genomics where they never would have known or cared before,” she says. “They’re talking about the output of sequencing as a result of things like CanCOGeN.”
COVID-19 has also given public-health labs the experience and equipment to do more sequencing, she points out. “We’re already working on our next genomics projects,” for example, focusing on tuberculosis and influenza. “It really has started us down the path of transforming a lot of work we’re doing over to genomics,” says Prystajecky, who predicts a continuing scale-up of sequencing across the public-health domain. “Genomics is now not just this geeky thing that we do on the side.”
Recruiting personnel trained in analytical fields like bioinformatics, data visualization and data integration will be an ongoing challenge, Prystajecky adds. “If genomics is here to stay, the analysis is here to stay as well.”
Poliquin notes that genomics is an esoteric topic on its face, and it took the COVID-19 crisis “to really help cement its power and its importance” in people’s minds. “There’s not a lot of silver linings to the pandemic,” he comments, but if it is helping to bring genomics “into people’s consciousness, and giving it the resources necessary to make a difference, well, I’ll take it as a silver lining.”
Key Takeaway: “Genomics” has entered the national vocabulary and we have a golden opportunity to step up science communications in Canada about the value and impact of genomics.
Preparing for the next pandemic
Annan says that future pandemic readiness is critical, which means continuing to collaborate across provincial boundaries and co-ordinate at the national level. “The lesson I hope we learn—and I’m worried that we won’t learn—is that we cannot take our eye off the ball when this thing passes.”
He feels that “COVID will be with us for a while yet,” and even when things get better in Canada, it won’t be extinguished everywhere. “There’s always the opportunity and possibility that a new variant pops up somewhere else in the world that suddenly is infectious in a way that we didn’t expect.” His bigger concern is that in 10 or 15 years, when we’re fighting deficits and dealing with other issues like climate change, there will be less interest in maintaining vigilance in this area. Meanwhile CanCOGeN offers guidance about the benefits of national co-ordination in tackling other things like cancer and the mental health crisis.
For Poliquin, a lesson of both SARS outbreaks is that an “agency that’s able to respond to pandemics has to be fed and kept healthy in between periods of crisis.” It’s necessary for policymakers to think about how Canada will remain ready for “routine pandemics,” he says. “How we keep the system primed and alive and healthy in those in-between moments is actually going to be the test that falls on all of us.”
He says that “the tools that we’ve developed for genomics for SARS-CoV-2 can be applied to all sorts of other crises we face.” These include understanding the transmission dynamics of tuberculosis in Canada’s northern regions and vector-borne illnesses like Lyme disease.
“The same way you can type COVID you can type Lyme. You can look at how it’s changing over time, how it’s spreading,” Poliquin explains. “There’s lots of other public-health questions that can be tackled with genomics. And we’ve built up an infrastructure that’s amongst the best in the world.”
Lopez-Correa notes that genomics technology can be used to monitor wastewater for the presence of viruses, even before people are sick in a community. “This will help us be more proactive and not reactive,” she says, and to understand viruses and how they behave from the beginning.
She feels that a major legacy of the pandemic is the tools and capabilities it has brought, which can continue to be put to good use to sequence HIV, salmonella and other pathogens. “Canada was a bit behind in our genomic surveillance in general. COVID was the opportunity to get us up to the right level.”
CanCOGeN is now in its final months as Genome Canada prepares to transition responsibility for COVID-19 genomic surveillance to the NML in the spring of 2022, as part of building sustainable genomics and viral-sequencing surveillance capacity.
Lopez-Correa this past summer was named chief scientific officer of Genome Canada, a position she will retain as she helps transform Genome Canada from a “program-driven” organization that finances applied research to a “mission-driven” one that uses genomics to address big global challenges like climate change and food security.
“The first big mission that we handled, of course, is COVID-19, with the CanCOGeN initiative,” she says. Genome Canada hopes to launch its next mission in January 2022, with an evolved focus and structure and a storehouse of lessons from the pandemic.
With Thanks to our Partners